# Data
library(sf)
library(readr)
library(tidyverse)
# Dates and Times
library(lubridate)
# graphs
library(ggplot2)
library(ggthemes)
library(gridExtra)
library(viridis)
library(ggimage)
library(grid)
library(plotly)
# Models
library(modelsummary)
library(broom)
library(gtools)9 Assessing Interventions
Since its outbreak in late 2019, COVID-19 affected millions of people worldwide. Daily case counts quickly became one of the main ways to track the pandemic and to debate the effects of lockdowns and other restrictions. These data helped policymakers monitor the evolution of the disease and make decisions about when to tighten or relax restrictions (Goodman-Bacon and Marcus 2020; Brodeur et al. 2021; Zhou and Kan 2021).
In this chapter, we use difference-in-differences (diff-in-diff or DiD), a widely used quasi-experimental design, to assess treatement, interventions or policy evaluation. We show both the promise and the limits of this approach. The aim is to estimate an effect and to learn how to judge whether a DiD design is credible. We begin with COVID case data and learn some practical lessons about the challenges on policy evaluation and identify different decision areas. We then switch to a small theoretical example to explain the mechanics of the method more clearly. This also prepares us for the assignment, where mobility data may offer a more plausible short-run outcome than recorded cases.
This chapter is based on:
Goodman-Bacon, Andrew, and Jan Marcus. “Using difference-in-differences to identify causal effects of COVID-19 policies.” (2020)
Andrew Heiss’ chapter on Difference-in-differences
Brodeur, Abel, et al. “COVID-19, lockdowns and well-being: Evidence from Google Trends.” Journal of public economics 193 (2021): 104346.
9.1 Dependencies
9.2 Data
First let’s import the Greater London COVID-19 data:
# import csv
covid_cases_london <- read.csv("data/longitudinal-2/covid_cases_london.csv", header = TRUE)
# check out the variables
colnames(covid_cases_london) [1] "areaType" "areaName"
[3] "areaCode" "date"
[5] "newCasesBySpecimenDate" "cumCasesBySpecimenDate"
[7] "newFirstEpisodesBySpecimenDate" "cumFirstEpisodesBySpecimenDate"
[9] "newReinfectionsBySpecimenDate" "cumReinfectionsBySpecimenDate" Then we do the same for the Lazio area data, which is the Region of the capital of Italy, Rome. We are choosing this region because it did not see sharp peaks in COVID-19 cases during the winter of 2020/2021.
# import csv
covid_cases_lazio <- read.csv("data/longitudinal-2/covid_cases_lazio.csv", header = TRUE)
# check out the variables
colnames(covid_cases_lazio) [1] "data" "stato"
[3] "codice_regione" "denominazione_regione"
[5] "codice_provincia" "denominazione_provincia"
[7] "sigla_provincia" "lat"
[9] "long" "totale_casi" First we need to clean up the data and rename some variables in both dataframes to have 4 variables:
date: year-month-daygeo: geographical regioncases: number of COVID-19 cases that dayarea: Lazio (Rome) or London
# Rename the variables in the Lombardia data frame
covid_cases_lazio_ren <- covid_cases_lazio %>%
rename(date = data , geo = denominazione_provincia, totalcases = totale_casi)
# Group the data by region and calculate total cases for each day (not cumulative cases)
covid_cases_lazio_daily <- covid_cases_lazio_ren %>%
group_by(geo) %>%
mutate(cases = totalcases - lag(totalcases, default = 0)) %>%
select(date, geo, cases) %>%
mutate(area = "Rome (Lazio)") %>%
filter(cases >= 0)
#df_milan <- covid_cases_lombardia_new %>%
# filter(geo == "Milano", new_cases >= 0)
# Rename the variables in the first data frame
covid_cases_london_ren <- covid_cases_london %>%
rename(date = date , geo = areaName, cases = newCasesBySpecimenDate) %>%
select(date, geo, cases) %>%
mutate(area = "London")
# Correct date format
covid_cases_london_ren$date <- as.Date(covid_cases_london_ren$date)
covid_cases_lazio_daily$date <- as.Date(covid_cases_lazio_daily$date)
# Append the renamed data frame to the second data frame
covid_combined <- rbind(covid_cases_london_ren, covid_cases_lazio_daily)
# Add a variable of log of cases
covid_combined <- covid_combined %>%
mutate(log_cases = log(cases))9.3 Data Exploration
Similarly to the previous chapter. Let’s start by eyeballing the data.
# Visualizing Cases in London
covid_cases_1 <- ggplot(data = covid_cases_london_ren, aes(x = date, y = cases, color=geo)) +
geom_line() +
scale_color_viridis(discrete = TRUE, option="magma") +
theme_minimal() +
theme(
legend.position = "bottom")
labs(
x = "",
y = "",
title = "Evolution in Covid-19 cases",
color = "Region"
) +
scale_x_date(date_breaks = "6 months")NULL covid_cases_1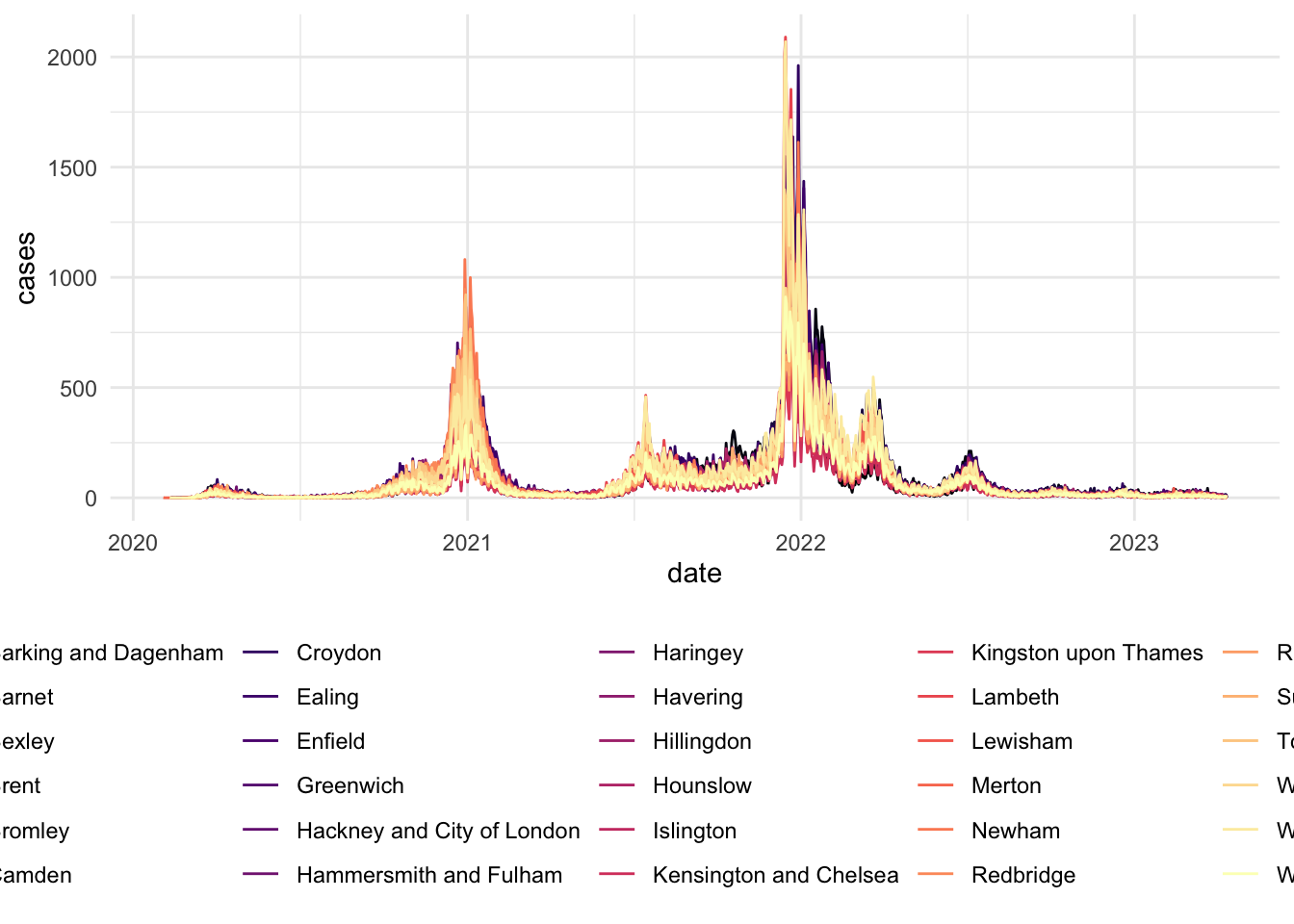
# Visualizing Cases in Lazio (Rome)
covid_cases_2 <- ggplot(data = covid_cases_lazio_daily, aes(x = date, y = cases, color=geo)) +
geom_line() +
scale_color_viridis(discrete = TRUE, option="magma") +
theme_minimal() +
theme(
legend.position = "bottom")
labs(
x = "",
y = "",
title = "Evolution in Covid-19 cases",
color = "Region"
) +
scale_x_date(date_breaks = "6 months")NULLcovid_cases_2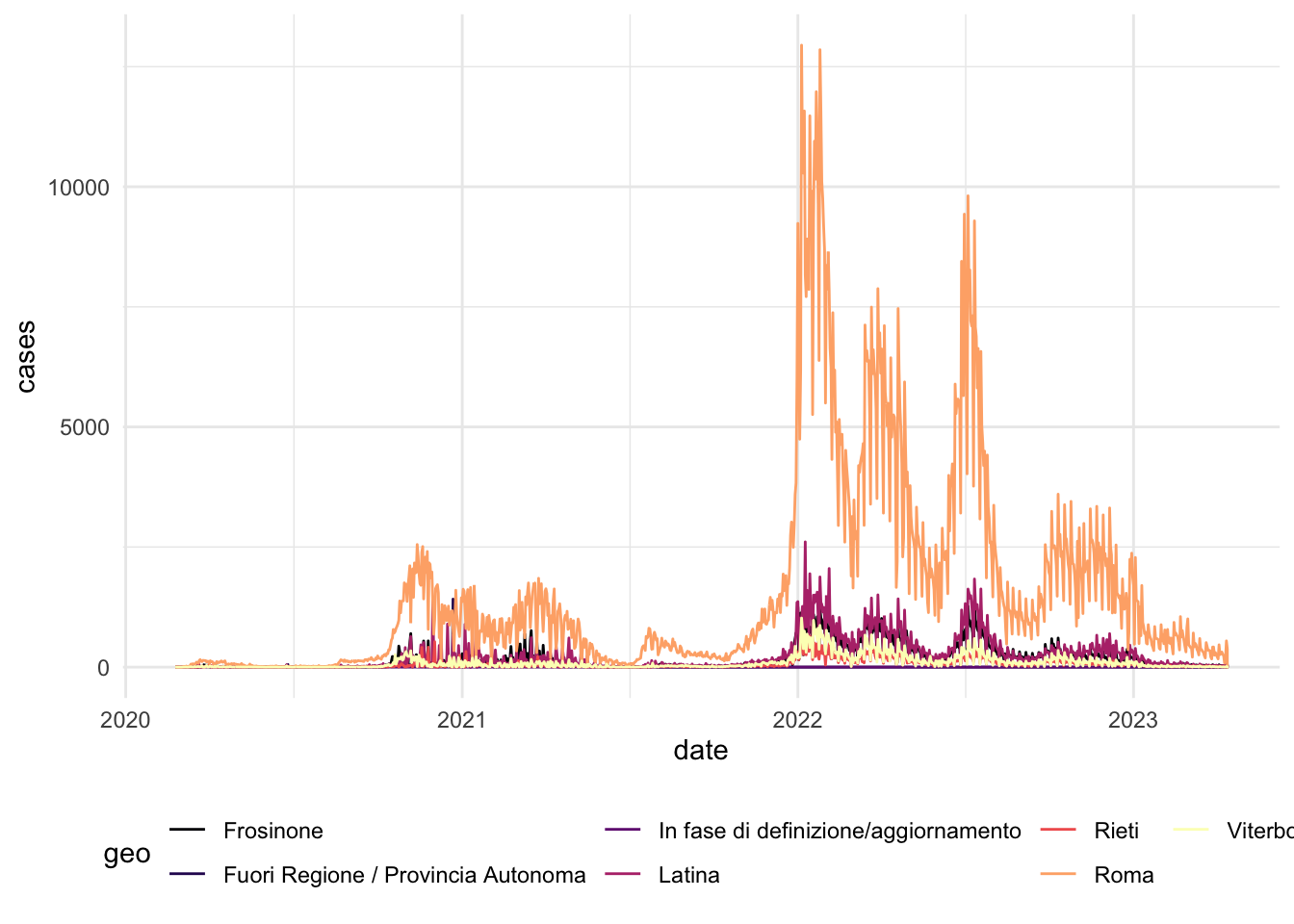
To identify the timing of lockdowns in these different areas and evolution of COVID-19 cases, we can overlap aggregage cases of both regions in a single plot.
# Aggregate the data by region for each day
covid_combined_agg <- aggregate(cases ~ area + date, data = covid_combined, FUN = sum)
# Visualizing aggregated
covid_cases_3 <- ggplot(data = covid_combined_agg, aes(x = date, y = cases, color=area)) +
geom_line() +
scale_color_manual(values=c("darkblue", "darkred")) + # set individual colors for the areas
theme_minimal() +
theme(
legend.position = "bottom"
) +
labs(
x = "",
y = "",
title = "Evolution in Covid-19 cases",
color = "Region"
) +
scale_x_date(date_breaks = "6 months")
covid_cases_3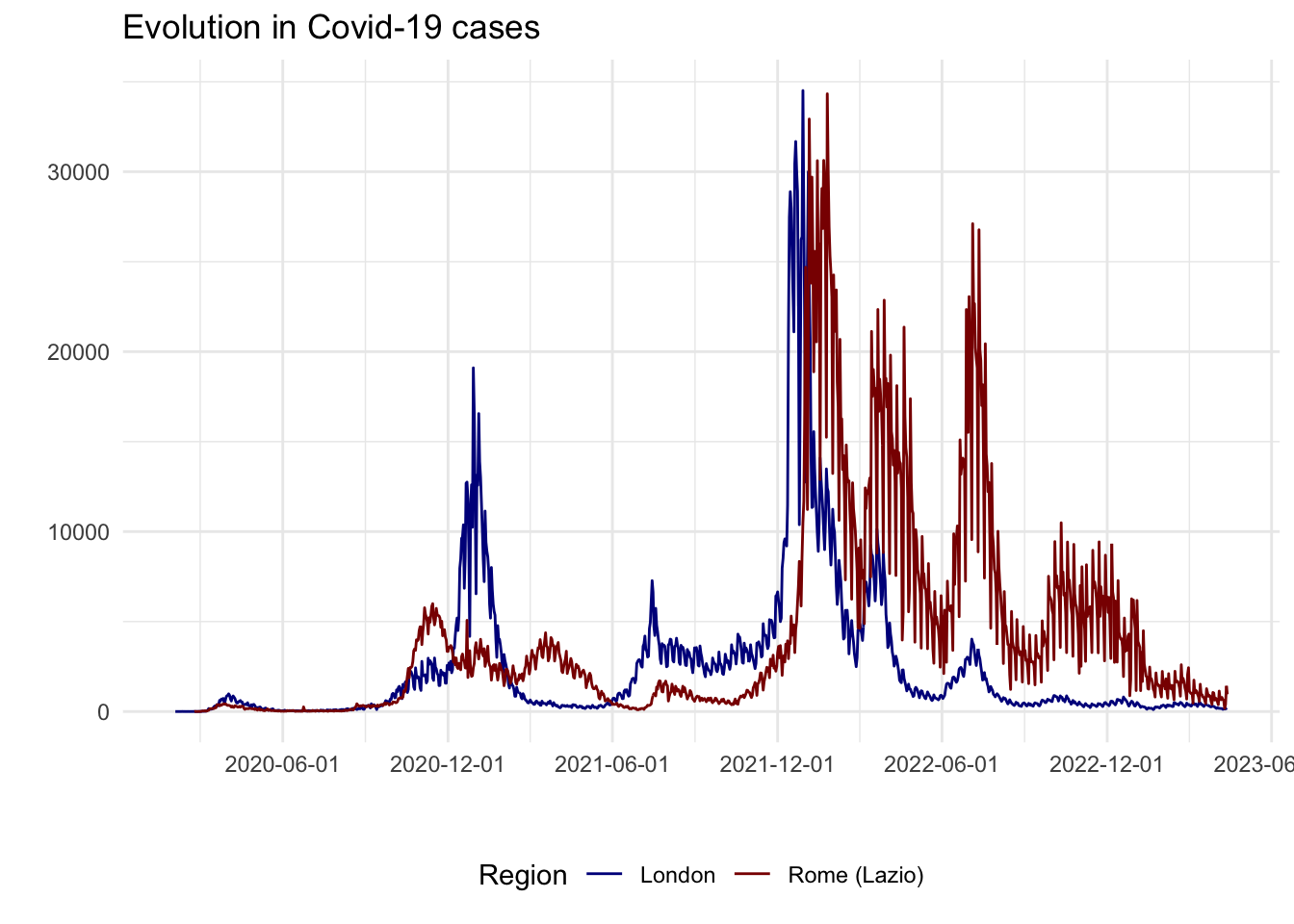
From an initial look at the data, the 2020/2021 winter period seems interesting as there is a high increase in London cases but not as much as a peak in Lazio cases. In fact, after a quick review of COVID-19 lockdowns, we found that:
- On the 5th of November 2020, the UK Prime Minister announced a second national lockdown, coming into force in England
- On 4 November 2020, Italian Prime Minister Conte also announced a new lockdown. Regions of the country were divided into three zones depending on the severity of the pandemic (not shown on the plot): red, orange and yellow zones. The Lazio region was a yellow zone for the duration of this second lockdown. In yellow zones, the only restrictions included compulsory closing for restaurant and bar activities at 6 PM, and online education for high schools only.
# Usual chart
covid_cases_4 <- ggplot(data = covid_combined_agg, aes(x = date, y = cases, color=area)) +
geom_line() +
scale_color_manual(values=c("darkblue", "darkred")) + # set individual colors for the areas
theme_minimal() +
theme(
legend.position = "bottom"
) +
labs(
x = "",
y = "",
title = "Evolution in Covid-19 cases",
color = "Region"
) +
scale_x_date(limit=c(as.Date("2020-08-01"), as.Date("2021-01-15"))) +
geom_vline(xintercept=as.numeric(as.Date("2020-11-05")), linetype="dashed") +
annotate("text", x=as.Date("2020-11-06"), y=25000, label="Lockdown",
color="black", fontface="bold", angle=0, hjust=0, vjust=0)
covid_cases_4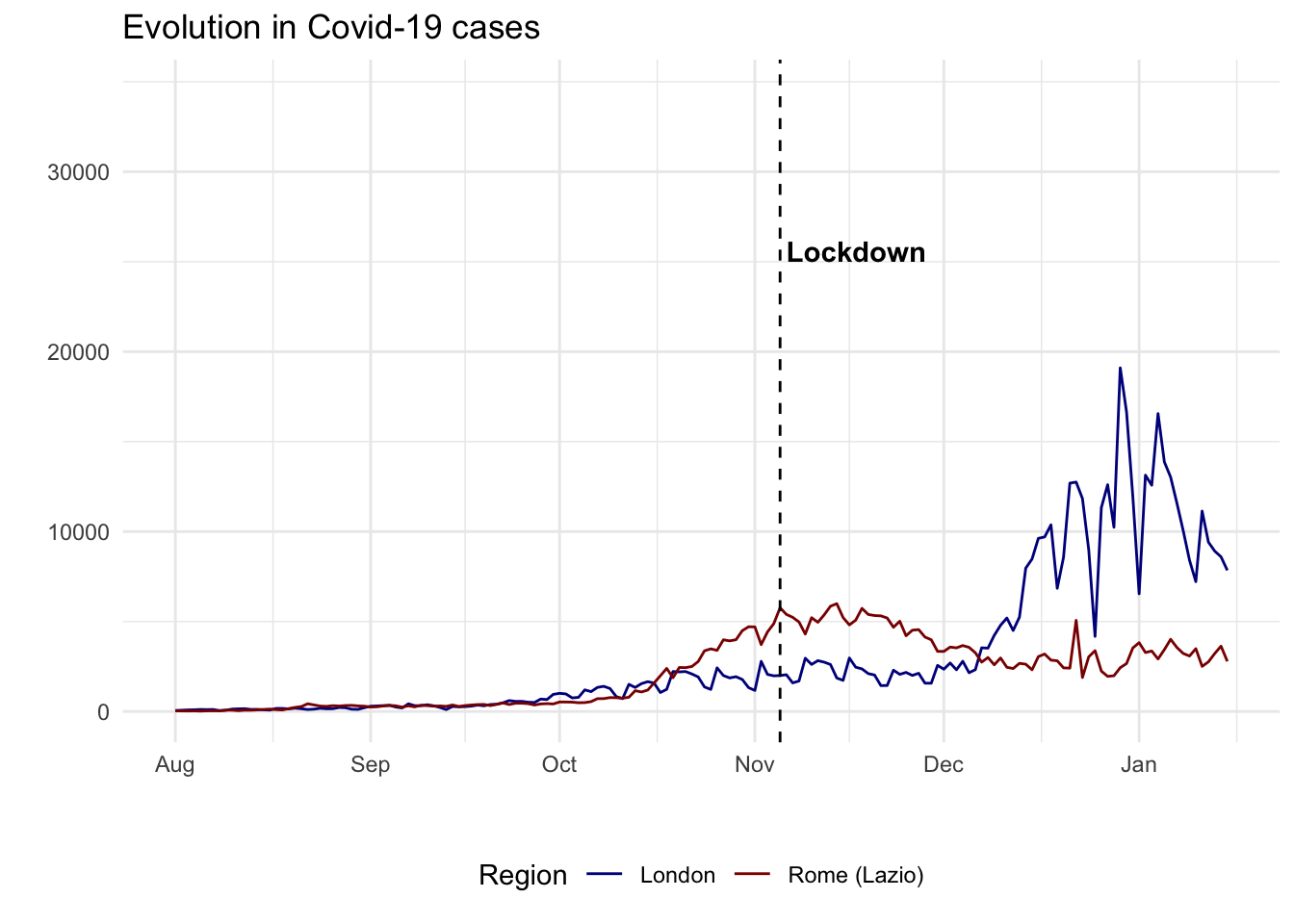
DiD is often conceptualised as a quasi-experiment. In social science, researchers often work with natural or quasi-experimental settings because randomized experiments are rarely possible. The key question to formulate such a quasi-experimental setting is to establish a credible comparison. As we will see below, a key component to this is establishing a reliable control or benchmark group of comparison.
9.4 Difference-in-Differences
What is difference in differences?
Difference-in-differences (DiD) is a way of estimating the effect of a policy or intervention when we cannot run a randomized experiment. Intuitively, we seek to compare how an outcome changes over time in a treated group with how the same outcome changes over time in a similar but untreated comparison group. A before-after comparison for the treated group alone is not enough, because many other things may be changing at the same time. A treated-versus-control comparison at one point in time is also not enough, because the two groups may already differ for other reasons. Difference-in-differences combines both comparisons and asks whether the treated group changed more, or less, than we would have expected given the change observed in the comparison group. Its key assumption is that, in the absence of treatment, the two groups would have followed parallel trends, so the untreated group provides a reasonable counterfactual for what would have happened to the treated group without the intervention.
Design choices
The DiD approach entails a number of critical choices. It includes a before-after comparison for a treatment and control group. In our example:
A
cross-sectional comparison(= compare a sample that was treated (London) to an untreated control group (Rome))A
before-after comparison(= compare treatment group with itself, before and after the treatment (5th of November))
The main and critical assumption of DiD is the parallel trends assumption. That means that, in the absence of the intervention, the treated and control groups would have followed similar trends over the period we analyse. The two groups do not need to have the same level, but they should evolve in a comparable way before treatment.
In practice, a good DiD design requires four choices: the treated group, the comparison group, the pre-treatment window, and the post-treatment window. For our example, we will see that making the right choices here may misguide our results and conclusions that we can draw from the data. Our comparison is imperfect because Lazio is not a pure untreated case. It also faced pandemic restrictions, even if they were milder than those in England. But, it is useful to illustrate the challenges of formulating a good quasi-experimental design for a credible DiD.
A useful first lesson from this chapter is that the time window matters. If we define the post period too broadly, we risk attributing later shocks to the policy of interest. Here that is especially important because London cases rose sharply again in December 2020 and January 2021.
# A wide window that mixes the November lockdown with the later winter surge
covid_combined_wide <- covid_combined %>%
filter(date >= "2020-09-01" & date <= "2021-01-01") %>%
mutate(after_5nov = ifelse(date >= "2020-11-05", 1, 0),
window = "Wide window: 1 Sep to 1 Jan")
# A tighter window focused on the weeks around the intervention
covid_combined_tight <- covid_combined %>%
filter(date >= "2020-10-15" & date <= "2020-11-30") %>%
mutate(after_5nov = ifelse(date >= "2020-11-05", 1, 0),
window = "Tighter window: 15 Oct to 30 Nov")
covid_designs <- bind_rows(covid_combined_wide, covid_combined_tight)We can now compare these two designs directly by plotting group means before and after 5 November.
design_plot_data <- covid_designs %>%
# Make these categories instead of 0/1 numbers so they look nicer in the plot
mutate(after_5nov = factor(after_5nov, labels = c("Pre 5 Nov 2020", "Post 5 Nov 2020"))) %>%
group_by(window, area, after_5nov) %>%
summarize(mean_cases = mean(cases),
se_cases = sd(cases) / sqrt(n()),
upper = mean_cases + (1.96 * se_cases),
lower = mean_cases + (-1.96 * se_cases),
.groups = "drop")
ggplot(design_plot_data, aes(x = after_5nov, y = mean_cases, color = area)) +
geom_pointrange(aes(ymin = lower, ymax = upper), size = 1) +
geom_line(aes(group = area)) +
scale_color_manual(values = c("darkblue", "darkred")) +
facet_wrap(vars(window))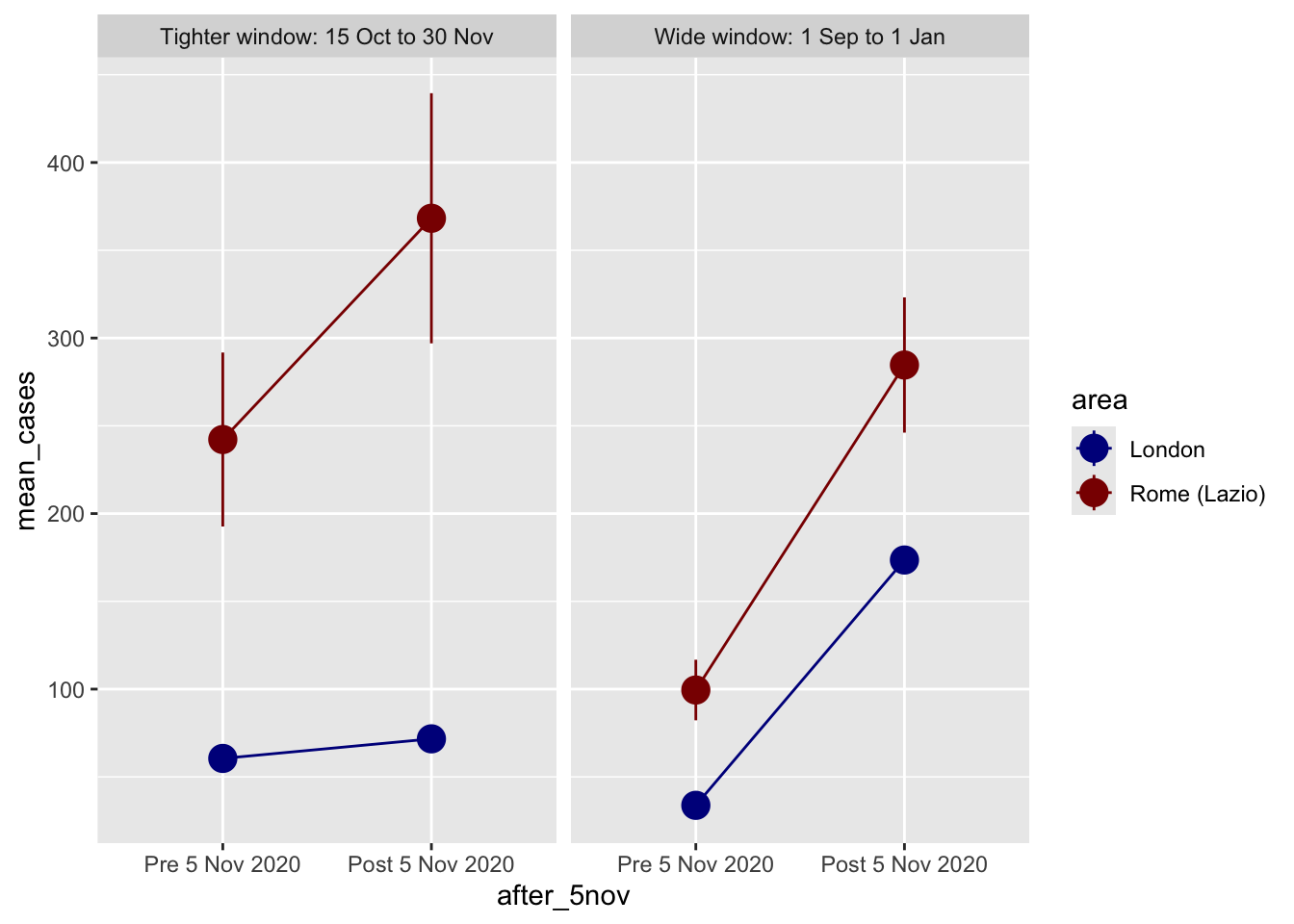
In our example, a shorter time window represents a better research design. It is a better reflection that the implementation of the lockdown in London contained the pre-treatment surge in Covid-19 cases. A wider window can be misleading as they suggest that the implementation of the lockdown did not have any impact. On the contrary, it is linked to a considerable increase in Covid-19. But from the time series plot above, we know that this actually reflects a surge in Covid-19 cases during December 2020 and January 2021.
9.4.1 Checking parallel trends
As highlighted above, a key issue to assess in the implementation of DiD is the compliance of the parallel trends assumption. In our example, this is responding to the question: whether London and Lazio were moving in similar ways before the intervention? The parallel trends assumption is about slopes, not levels. One group can start higher than the other, but the pre-treatment trajectories should be broadly similar.
A practical way to assess this is:
- Plot only the pre-treatment period.
- Look for similar direction and pace of change.
- Use domain knowledge to ask whether other shocks or policies were affecting the groups differently.
- If useful, run a simple pre-trend regression and check whether the treated and control groups have different pre-treatment slopes.
pre_period <- covid_combined_tight %>%
group_by(area, date) %>%
summarize(cases = sum(cases), .groups = "drop") %>%
filter(date < as.Date("2020-11-05")) %>%
mutate(day = as.numeric(date - min(date)),
treat_london = ifelse(area == "London", 1, 0))
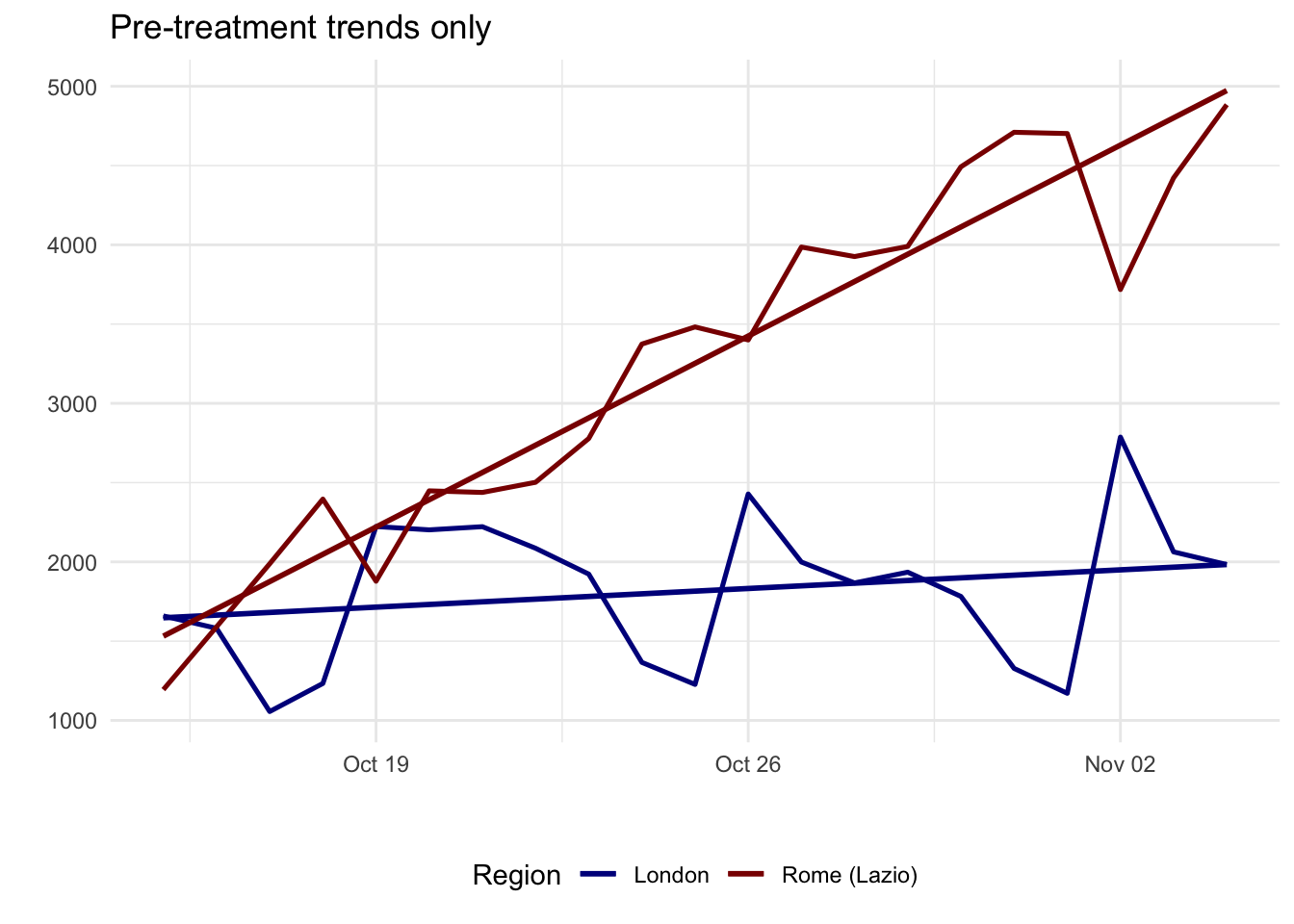
ggplot(pre_period, aes(x = date, y = cases, color = area)) +
geom_line(aes(group = area), linewidth = 0.9) +
geom_smooth(method = "lm", formula = y ~ x, se = FALSE, linewidth = 1) +
scale_color_manual(values = c("darkblue", "darkred")) +
theme_minimal() +
theme(
legend.position = "bottom"
) +
labs(
x = "",
y = "",
title = "Pre-treatment trends only",
color = "Region"
)
pretrend_model <- lm(cases ~ treat_london + day + treat_london:day,
data = pre_period)
tidy(pretrend_model) %>%
filter(term == "treat_london:day")# A tibble: 1 × 5
term estimate std.error statistic p.value
<chr> <dbl> <dbl> <dbl> <dbl>
1 treat_london:day -155. 20.5 -7.57 0.00000000424In this case, the pre-treatment slopes are different, so the London-Lazio comparison is a weak comparison. The narrower window fixes one design problem, but it does not fully offer a credible quasi-experimental design. A pre-trend test like this is only a diagnostic, not proof. It is a useful warning sign when the interaction term is large and clearly different from zero.
9.4.2 What if parallel trends fail?
If the parallel trends assumption does not look plausible, do not force the method. A sensible sequence is:
- Redefine the time window so the pre and post periods match the timing of the policy more closely.
- Allow for a plausible lag if the outcome reacts with delay, as COVID case data often do.
- Choose a better comparison group with a more similar pre-treatment trend and policy context.
- Change the outcome to something that responds more immediately to the intervention, such as mobility or another behavioural measure.
- If none of these improve the design, present the comparison as descriptive evidence and use another research design for causal claims.
This is the point at which the London-Lazio example becomes most useful. It shows how DiD helps us diagnose a weak comparison instead of blindly producing a coefficient. To explain the mechanics of DiD more clearly, we now switch to a small theoretical example where the identifying logic is closer to what we want.
# import csv
covid_perfect_example <- read_csv("data/longitudinal-2/example_covid.csv")
# label pre/post labels
covid_perfect_example <- covid_perfect_example %>%
mutate(after_5nov = factor(after_5nov, labels = c("Before 5 November 2020", "After 5 November 2020")))
# plot
ggplot(covid_perfect_example, aes(x = after_5nov, y = mean_cases, color = area)) +
geom_pointrange(aes(ymin = lower, ymax = upper), size = 1) +
geom_line(aes(group = area)) +
scale_color_manual(values = c("darkblue", "darkred"))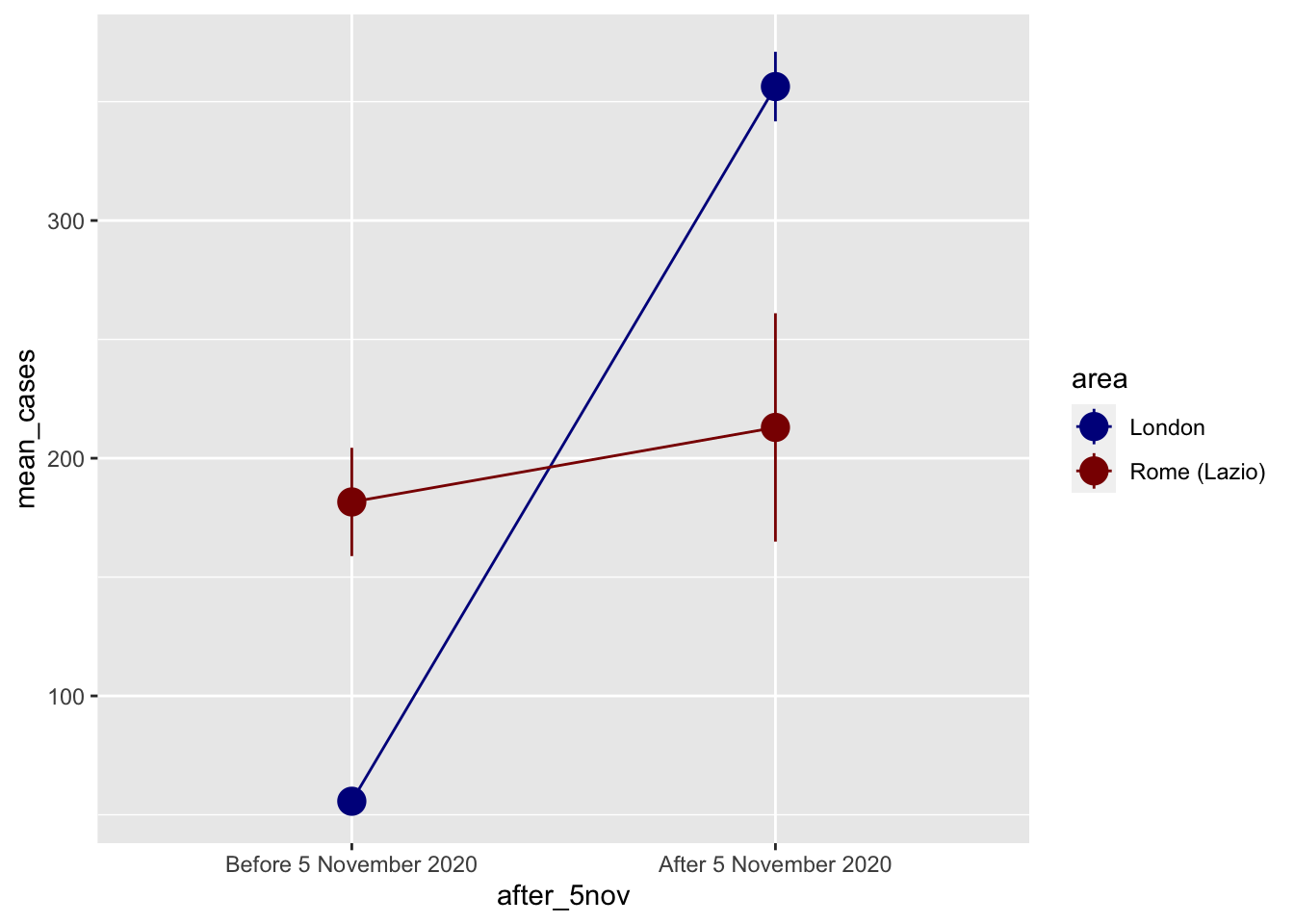
Difference in Difference by hand
We can find the exact difference by filling out the 2x2 before/after treatment/control table:
| Before | After | Difference | |
|---|---|---|---|
| Treatment | A | B | B - A |
| Control | C | D | D - C |
| Difference | C - A | D - B | (D − C) − (B − A) |
We can pull each of these numbers out of the table:
before_treatment <- covid_perfect_example %>%
filter(after_5nov == "Before 5 November 2020", area == "London") %>%
pull(mean_cases)
before_control <- covid_perfect_example %>%
filter(after_5nov == "Before 5 November 2020", area == "Lazio") %>%
pull(mean_cases)
after_treatment <- covid_perfect_example %>%
filter(after_5nov == "After 5 November 2020", area == "London") %>%
pull(mean_cases)
after_control <- covid_perfect_example %>%
filter(after_5nov == "After 5 November 2020", area == "Lazio") %>%
pull(mean_cases)
diff_treatment_before_after <- after_treatment - before_treatment
diff_treatment_before_after[1] 156.35diff_control_before_after <- after_control - before_control
diff_control_before_after[1] 24.35387diff_diff <- diff_treatment_before_after - diff_control_before_after
diff_diff[1] 131.9961In this theoretical example, the DiD estimate is 131.99. This is the gap between the treated group’s observed post-treatment outcome and the counterfactual post-treatment outcome implied by the control trend.
We can visualise this with a bit of extra code:
ggplot(covid_perfect_example, aes(x = after_5nov, y = mean_cases, color = area)) +
geom_point() +
#geom_pointrange(aes(ymin = lower, ymax = upper), size = 1) +
#geom_line(aes(group = area)) +
geom_line(aes(group = as.factor(area))) +
scale_color_manual(values = c("darkblue", "darkred")) +
# If you use these lines you'll get some extra annotation lines and
# labels. The annotate() function lets you put stuff on a ggplot that's not
# part of a dataset. Normally with geom_line, geom_point, etc., you have to
# plot data that is in columns. With annotate() you can specify your own x and
# y values.
annotate(geom = "segment", x = "Before 5 November 2020", xend = "After 5 November 2020",
y = before_treatment, yend = after_treatment - diff_diff,
linetype = "dashed", color = "grey50") +
annotate(geom = "segment", x = "After 5 November 2020", xend = "After 5 November 2020",
y = after_treatment, yend = after_treatment - diff_diff,
linetype = "dotted", color = "blue") +
annotate(geom = "label", x = "After 5 November 2020", y = after_treatment - (diff_diff / 2),
label = "Program effect", size = 3)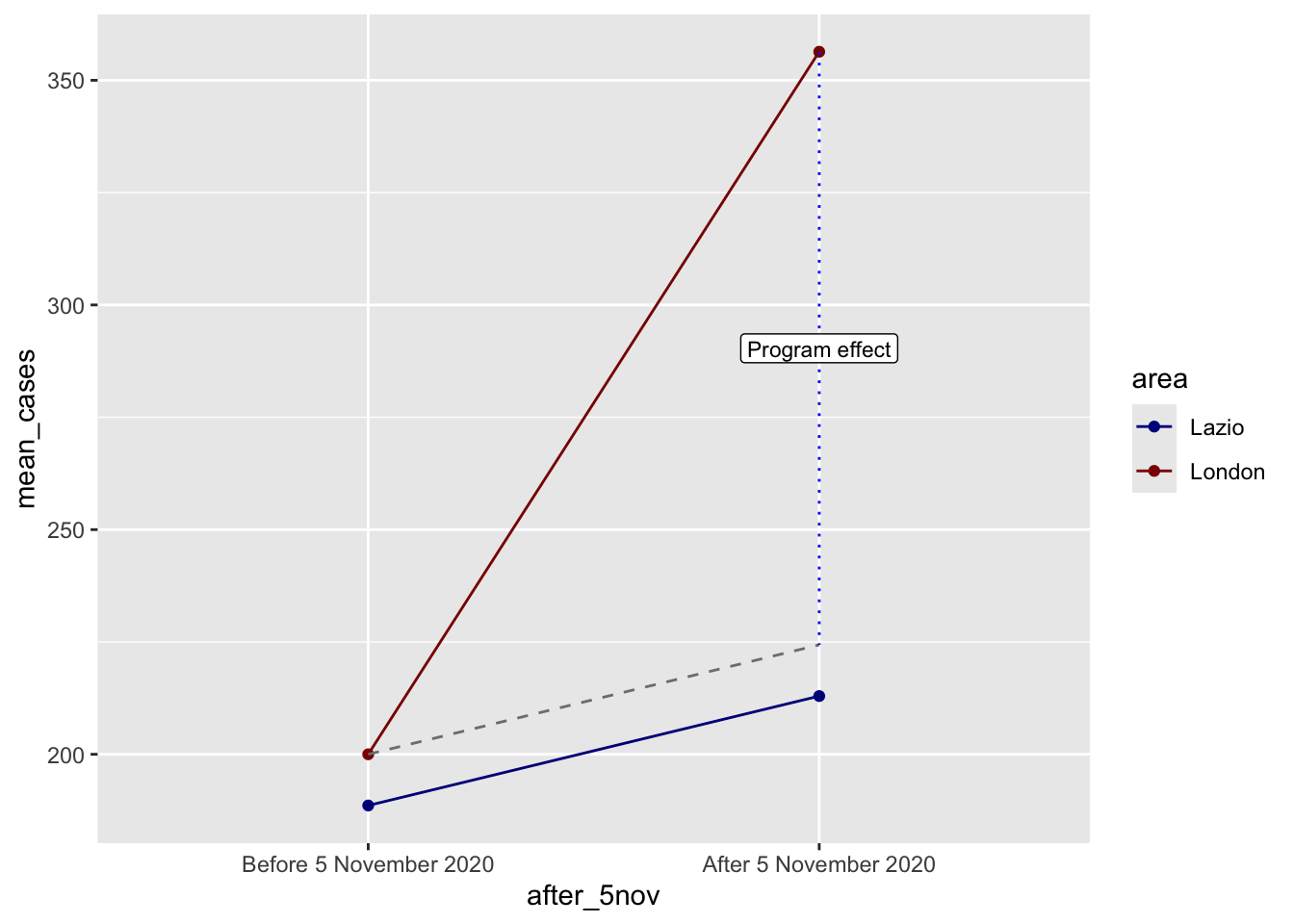
In this scenario, we have a credible quasi-experimental design. DiD shows the importance of establishing credible comparison groups and time frame to assess the influence of interventions. As with COVID data, a bad experimental design may produce misleading results because of it may capture the compounding effects of policy bundles, reverse causality, anticipation effects, changes in testing and data collection, and variation in policy timing (Goodman-Bacon and Marcus 2020). These problems are especially serious when the outcome is recorded cases.
Difference-in-Difference with regression
In practice, our control and treated groups tend to differ in systematic ways. Often, then, we want to account for these systematic differences by running DiD analysis in a regression framework. To this end, we first code London as the treated group and the post-5 November period as the post-treatment period:
covid_perfect_reg <- covid_perfect_example %>%
mutate(
treat = ifelse(area == "London", 1, 0),
post = ifelse(after_5nov == "After 5 November 2020", 1, 0)
)We can now build the regression in four steps:
mean_cases ~ treatThis is only a cross-sectional comparison. It tells us whether London and Lazio differ on average, but it ignores time completely.mean_cases ~ postThis is only a before-after comparison. It tells us whether average cases are different after 5 November, but it ignores whether the change happened in the treated or control group.mean_cases ~ treat + postThis controls for baseline group differences and for a common time shift, but it still does not isolate the treatment effect.mean_cases ~ treat * postThis adds the interaction term. That interaction is the DiD estimate.
model_treat <- lm(mean_cases ~ treat, data = covid_perfect_reg)
model_post <- lm(mean_cases ~ post, data = covid_perfect_reg)
model_additive <- lm(mean_cases ~ treat + post, data = covid_perfect_reg)
model_did <- lm(mean_cases ~ treat * post, data = covid_perfect_reg)
modelsummary(
list(
"Treat only" = model_treat,
"Post only" = model_post,
"Treat + post" = model_additive,
"Treat * post" = model_did
),
coef_map = c(
"(Intercept)" = "Control group, before",
"treat" = "Treat: London",
"post" = "Post period",
"treat:post" = "DiD estimate"
),
estimate = "{estimate}",
statistic = NULL,
gof_omit = ".*"
)| Treat only | Post only | Treat + post | Treat * post | |
|---|---|---|---|---|
| Control group, before | 200.781 | 194.302 | 155.605 | 188.604 |
| Treat: London | 77.394 | 77.394 | 11.396 | |
| Post period | 90.352 | 90.352 | 24.354 | |
| DiD estimate | 131.996 |
Because this theoretical dataset contains only the four DiD cells, we focus on the coefficients rather than on standard errors or significance tests. The point here is interpretation:
- In model 1, the
treatcoefficient captures the average London-Lazio difference, but that mixes pre-treatment and post-treatment differences together. - In model 2, the
postcoefficient captures the average change after 5 November, but that mixes treated and control areas together. - In model 3, the
treatandpostcoefficients separate group differences from time differences, but they still assume the treatment effect is zero for both groups. - In model 4, the interaction term
treat:postis the extra post-treatment change in London relative to Lazio. This is the DiD estimate, and it matches the quantity we calculated by hand above.
The final regression is therefore:
\(Y_{gt} = \beta_0 + \beta_1 Treat_g + \beta_2 Post_t + \beta_3 (Treat_g \times Post_t) + \epsilon_{gt}\)
We could then expand this regression by including relevant covariates of interest. Feel free to expand this basic structure adding covariates or identifying a more credible, truly untreated benchmark group. DiD works best when: (1) a policy changes for one group but not the group used for comparison; (2) the timing of the intervention is clear, and (3) pre-treatment trends are reasonably similar. Well-known applications include:
- labour economics: comparing employment in New Jersey and Pennsylvania after a minimum wage change (Card and Krueger 1993).
- public health: comparing places that expanded Medicaid (Wiggins, Karaye, and Horney 2020) or adopted smoking bans with similar places that did not (Fu et al. 2024).
- urban and environmental policy: comparing cities that introduced congestion charging (Li, Graham, and Majumdar 2012) or low-emission zones with cities that did not (Sarmiento, Wägner, and Zaklan 2023)
- migration: assessing the effects of visa policy changes (Faggian, Corcoran, and Rowe 2015)
9.5 Questions
For the assignment, you will continue to use Google Mobility data for the UK for 2021. For details on the timeline you can have a look here. You will need to do a bit of digging on when lockdowns or other COVID-19 related shocks happened in 2021 to set up a DiD strategy. Have a look at Brodeur et al. (2021) to get some inspiration. They used Google Trends data to test whether COVID-19 and the associated lockdowns implemented in Europe and America led to changes in well-being.
Start by loading both the csv
mobility_gb <- read.csv("data/longitudinal-1/2021_GB_Region_Mobility_Report.csv", header = TRUE)Visualize the data with
ggplotand identify a COVID-19 intervention, a treated group, a comparison group, and a before/after window. Examples of these interventions could be a regional lockdown, school closures, travel restrictions, or vaccine rollouts. Generate a cleanggplotwhich indicates which intervention you are going to examine and explain briefly why this could be a useful DiD design.Plot the pre-treatment period only and assess whether the parallel trends assumption looks plausible. If it does not, redesign your setup once by changing the control group, the time window, or the outcome.
Define and estimate a DiD regression for your final design. What does the interaction term suggest? Would you treat it as a credible causal effect? Discuss the reasons why or why not.
Discuss how the unique dynamics of COVID-19 and the possibility of policies having differential effects over time complicate the interpretation of your results. If your design remains weak, explain what alternative outcome or design you would choose next.
Analyse and discuss what insights you obtain into people’s changes in behaviour during the pandemic in response to an intervention.